Saturday, January 11, 2020
New deadline for Yahoo Groups data request
|
Add to: CiteUlike | Connotea | Del.icio.us | Digg | Furl | Newsvine | Reddit | Yahoo
Wednesday, December 11, 2019
Next Steps: The Evolution of Yahoo Groups (Final Notification)
|
Add to: CiteUlike | Connotea | Del.icio.us | Digg | Furl | Newsvine | Reddit | Yahoo
Friday, November 08, 2019
Yahoo Groups - Upcoming Product Changes to Yahoo Groups
|
Add to: CiteUlike | Connotea | Del.icio.us | Digg | Furl | Newsvine | Reddit | Yahoo
Tuesday, January 16, 2007
Body Size, Performance and Fitness in Galapagos Marine Iguanas
Integrative and Comparative Biology 2003 43(3):376-386; doi:10.1093/icb/43.3.376
Martin Wikelski and L. Michael Romero
Complex organismal traits such as body size are influenced by innumerable selective pressures, making the prediction of evolutionary trajectories for those traits difficult. A potentially powerful way to predict fitness in natural systems is to study the composite response of individuals in terms of performance measures, such as foraging or reproductive performance. Once key performance measures are identified in this top-down approach, we can determine the underlying physiological mechanisms and gain predictive power over long-term evolutionary processes. Here we use marine iguanas as a model system where body size differs by more than one order of magnitude between island populations. We identified foraging efficiency as the main performance measure that constrains body size. Mechanistically, foraging performance is determined by food pasture height and the thermal environment, influencing intake and digestion. Stress hormones may be a flexible way of influencing an individual's response to low-food situations that may be caused by high population density, famines, or anthropogenic disturbances like oil spills. Reproductive performance, on the other hand, increases with body size and is mediated by higher survival of larger hatchlings from larger females and increased mating success of larger males. Reproductive performance of males may be adjusted via plastic hormonal feedback mechanisms that allow individuals to assess their social rank annually within the current population size structure. When integrated, these data suggest that reproductive performance favors increased body size (influenced by reproductive hormones), with an overall limit imposed by foraging performance (influenced by stress hormones). Based on our mechanistic understanding of individual performances we predicted an evolutionary increase in maximum body size caused by global warming trends. We support this prediction using specimens collected during 1905. We also show in a common-garden experiment that body size may have a genetic component in iguanids. This 'performance paradigm' allows predictions about adaptive evolution in natural populations. [Galapagos Islands]
-------
Recent post: Why are lions not as big as elephants?
Technorati: integrative, comparative, biology, complex, organismal, traits, body, size, evolutionary, fitness, natural, systems, performance, foraging, reproductive, marine, iguanas, island, stress, hormones, global warming, trends, specimens, genetic, adaptive, evolution, galapagos, islands, lions, elephants
Add to: CiteUlike | Connotea | Del.icio.us | Digg | Furl | Newsvine | Reddit | Yahoo
Wednesday, December 13, 2006
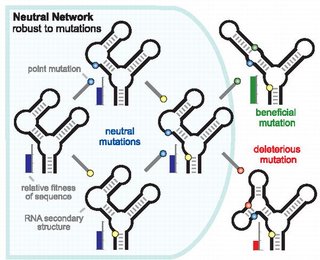
Balancing Robustness and Evolvability
From PloS Biology:
Balancing Robustness and Evolvability
Richard E. Lenski et al.
One of the most important features of biology is the ability of organisms to persist in the face of changing conditions. Consider the remarkable fact that every organism alive today is the product of billions of generations in which its progenitors, without fail, managed to produce progeny that survived to reproduce. To achieve this consistency, organisms must have a balance between robustness and evolvability, that is, between resisting and allowing change in their own internal states [1 - 3]. Moreover, they must achieve this balance on multiple time scales, including physiological responses to changes over an individual life and evolutionary responses, in which a population of genomes continually updates its encoded information about past environments and how future generations should respond given that record.
Examples of robust biological systems are found at many scales, from biochemical to ecological. At each scale, robustness may reflect the properties of individual elements or, alternatively, the dynamic feedbacks between interacting elements. The expression of some metabolic function, for example, may be robust in the face of temperature change, because an enzyme maintains its shape and specificity across a range of temperatures or because an interconnected network of reactions sustains the supply of product, even when some enzyme fails. A genome may be robust because it encodes proofreading and repair systems that reduce replication errors or because it is organized such that many mutations have little effect on its phenotype. An ecosystem might be robust if it resists the extinction of some keystone species or, if extinction does occur, because surviving species can compensate over physiological, demographic, or evolutionary time scales.
One important question is whether there exists a single unifying mathematical framework that can encompass such diverse examples of biological robustness. Might new insights come from such a conceptual unification, or will future understanding require detailed analyses of specific cases? Across the different scales, recurring mechanisms for achieving robustness - including redundancy of component parts and negative feedbacks - might serve as organizing principles. Yet, similarities in mechanism could mask important differences in the evolutionary origins of those mechanisms. At the level of genes in genomes or of cells in multicellular organisms, it is reasonable to suggest that redundancy evolved by natural selection to maintain some functional capacity in the face of perturbation [4]. But whereas species redundancy could also be critical for robustness of ecosystem functions, differences in redundancy might be an emergent property rather than an ecosystem-level adaptation, because selection generally acts at lower levels (but see [5] for another view).
Continued at "Balancing Robustness and Evolvability" [A modified version of this post (with background info) will be posted to the "General Evolution News" category]
Technorati: balancing, robustness, evolvability, biology, organisms, conditions, generations, progeny, genome, repair, systems, framework, genes, evolutionary, origins, multicellular, demographic, physiological, ecosytem, redundancy, richard, e, lenski, adaptation, phenotype, natural selection, emergent, property, level, keystone, species, extinction
Add to: CiteUlike | Connotea | Del.icio.us | Digg | Furl | Newsvine | Reddit | Yahoo
Sunday, November 26, 2006
Genomic Imprinting in Mammals: Emerging Themes and Established Theories
[This post also appears in the General Evolution News category]
An open access/free review paper from PLoS Genetics:
Genomic Imprinting in Mammals: Emerging Themes and Established Theories
Andrew J. Wood, Rebecca J. Oakey
The epigenetic events that occur during the development of the mammalian embryo are essential for correct gene expression and cell-lineage determination. Imprinted genes are expressed from only one parental allele due to differential epigenetic marks that are established during gametogenesis. Several theories have been proposed to explain the role that genomic imprinting has played over the course of mammalian evolution, but at present it is not clear if a single hypothesis can fully account for the diversity of roles that imprinted genes play. In this review, we discuss efforts to define the extent of imprinting in the mouse genome, and suggest that different imprinted loci may have been wrought by distinct evolutionary forces. We focus on a group of small imprinted domains, which consist of paternally expressed genes embedded within introns of multiexonic transcripts, to discuss the evolution of imprinting at these loci.
Introduction
The process of sexual reproduction dictates that mammals inherit two copies of every gene, one from the mother and one from the father. At most loci, both alleles are actively transcribed and functionally equivalent. Imprinted genes represent an exception to this rule, as the transcriptional activity of each allele is determined by the gender of the parental germ line to which it was most recently exposed. This parental legacy is initiated by epigenetic modifications such as DNA methylation, which is established in the parental germ line and maintained throughout somatic development in the offspring. Individual germ-line marks can control the allele-specific silencing or activation of multiple neighbouring genes, which leads in many instances to clusters of imprinted transcripts. Such loci represent an attractive paradigm for the study of epigenetic transcriptional regulation, as both the active and silent allele are present in the same cell nucleus, and therefore potentially exposed to the same trans-acting regulatory factors. Epigenetic abnormalities at imprinted loci have been observed in cloned mammals [1], and their disruption has been reported in a number of human developmental disorders and cancers [2].
Defining the Extent of Imprinting
Since the identification of the first autosomal imprinted genes in the early 1990s [3–5], much speculation has surrounded the question of how many exist. Attempts to count the exact number have been complicated by difficulties in defining exactly what constitutes a gene, as in several cases multiple functional components are derived from a single core of genetic information [6]. A recent census identified 96 imprinted functional components (54 maternally expressed, 42 paternally expressed) arising from 71 transcriptional units [7], and the relevant literature is summarised on the Harwell and University of Otago online databases [8,9].
A number of different approaches have been employed to define the extent of imprinting in the mouse genome. Mouse stocks carrying translocation chromosomes were used to define chromosomal regions that show parent-of-origin effects on phenotype when uniparentally inherited, and at least 13 distinct regions on eight chromosomes have been identified by this approach (C. V. Beechey, personal communication; [8]). The phenotypes range from early embryonic lethality to postnatal effects on growth and development, and are likely to result from the misexpression of imprinted genes situated within the uniparentally duplicated region [10]. The subsequent identification of imprinted genes on chromosomes without obvious uniparental effects [11-13] suggests that imprinting may be more widespread than initially thought, and not limited to genes that are vital for development. This conclusion is supported by the involvement of imprinted genes in behavioural traits in the mouse [14,15].
Continued at "Genomic Imprinting in Mammals: Emerging Themes and Established Theories"
-------
Featured Book: "Chromatin and Gene Regulation: Mechanisms in Epigenetics" by Bryan M. Turner (Amazon Astore UK | US)
Books on Epigenetics from the Science and Evolution Bookshop: UK | US
Technorati: open access, plos, genetics, imprinting, mammals, epigenetic, embryo, cell, marks, genomic, evolutionary, forces, domains, genes, evolution, dna, methylation, genetic, genome, mouse, chromosomes, epigenetics, science, regions
Add to: CiteUlike | Connotea | Del.icio.us | Digg | Furl | Newsvine | Reddit | Yahoo
Tuesday, November 14, 2006
Epigenetics: Mother's Diet during Pregnancy can affect Grandchildren
[This post also appears in the General Evolution News category]
Oakland, California: A new study by scientists at Children's Hospital Oakland Research Institute (CHORI) is the first to show that a mother's diet during pregnancy influences the health of her grandchildren by changing the behavior of a specific gene. The study was conducted using mice of an unique strain called 'viable yellow agouti' also known as A-vy in scientific terms. These mice possesss a gene that influences the color of their coats as well as their tendency to become obese and develop diabetes and cancer. The new research shows that the diet consumed by a pregnant Avy mouse affects the health of not only her pups, but also their pups - her grandchildren.
The study will be published in the November issue of the Proceedings of the National Academy of Sciences (PNAS) and was conducted by CHORI Scientist David Martin, M.D., and Assistant Scientist Kenneth Beckman, Ph.D., in collaboration with Drs. Jennifer Cropley and Catherine Suter from the Victor Chang Heart Institute in Sydney, Australia. In their experiments, the scientists fed some Avy mice a standard lab diet based on common foods consumed by humans. Other mice were fed this same diet supplemented with common nutritional supplements including folate, choline, betaine, vitamin B12, zinc and methionine.
The supplements were fed to the mice for a week during mid-pregnancy. The offspring were examined for their coat color, and female offspring were themselves mated again (without a supplemented diet) to produce a third generation of 'grandchildren.' The results showed that the supplements changed the behavior of the agouti gene in the first generation of pups, shifting their coats towards a brown color, and had the same effect on pups born in the next generation to mice that were not exposed to the supplemented diet.
Continued at "Epigenetics: Mother's Diet during Pregnancy can affect Grandchildren" [Evolution, Science]
-------
Based on the PNAS paper "Germ-line epigenetic modification of the murine A-vy allele by nutritional supplementation" (Abstract)
See "Cardiovascular and diabetes mortality determined by nutrition" (very relevant - don't be misled by the title!)
And "New theory of environmental inheritance ('05 Press Release)"
And "Epigenetics: Parentage has effects outside the genome"
Books on Epigenetics from the Science and Evolution Bookshop: UK | US
Epigenetic books from the Science and Evolution Bookshop: UK | US
Technorati: california, oakland, children, hospital, research, institute, chori, mother, diet, pregnancy, health, grandchildren, behavior, gene, mice, agouti, color, obese, diabetes, cancer, pups, mouse, pnas, academy, sciences, national, heart, sydney, australia, epigenetics, evolution, science, epigenetic, genome, enviromental, inheritance
Add to: CiteUlike | Connotea | Del.icio.us | Digg | Furl | Newsvine | Reddit | Yahoo